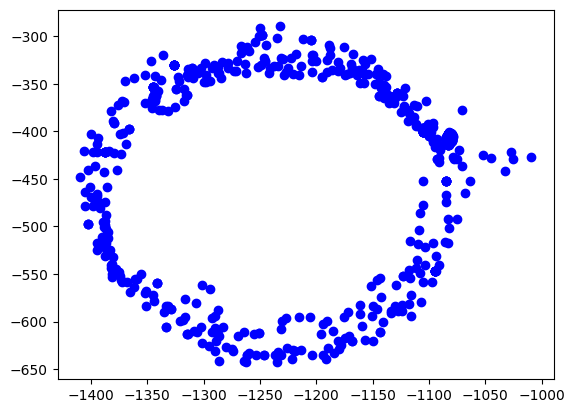
python matplotlib 如何使x,y轴的单位长度相等
在使用Python的matplotlib库绘制图形时,我们常常需要控制坐标轴的单位长度。
当x和y轴的比例不同,图形可能会被拉伸或者压缩,从而失真。本文将介绍如何通过设置坐标轴的纵横比例,
使得x和y轴的单位长度相等。
Matplotlib是一个功能强大的Python绘图库,可用于创建各种类型的静态、动态和交互式图形。
它提供了许多选项和配置,以便用户可以自定义他们的绘图。其中一个重要的功能就是控制坐标轴的纵横比例。
在Matplotlib中,我们可以使用axis()函数来设置坐标轴的范围和纵横比例。
具体来说,axis()函数有四个参数:[xmin, xmax, ymin, ymax]。这些参数控制了x和y轴的范围。
如果我们只提供前两个参数,则Matplotlib将使用默认值。
接下来,我们可以使用aspect参数来控制坐标轴的纵横比例。该aspect参数可以是一个浮点数或字符串(如"equal")。
如果我们将aspect参数设置为"equal",则x和y轴的单位长度将相等。
否则,我们可以计算出x和y轴的比例,并将其作为浮点数提供给aspect参数。
下面,我们通过一个示例来演示如何使用Matplotlib设置坐标轴的纵横比例。
首先,我们需要导入Matplotlib库,并创建一个Figure对象和一个Axes对象。
然后,我们使用plot()函数生成一些随机数据并将其绘制在图形上。
import matplotlib.pyplot as pltimport numpy as np# 创建Figure对象和Axes对象fig, ax = plt.subplots()# 生成随机数据x = np.arange(0, 10)y = np.random.rand(10)# 绘制线条ax.plot(x, y)现在,我们将使用axis()方法控制坐标轴的范围和纵横比例。
在这里,我们将指定x轴的范围为[0, 10],y轴的范围为[0, 1],并将aspect参数设置为"equal":
# 设置坐标轴范围和aspect参数ax.axis([0, 10, 0, 1])
ax.set_aspect("equal")
最后,我们通过show()方法显示图形:
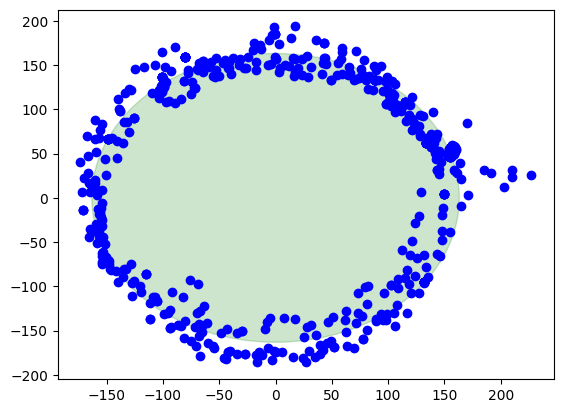
plt.show()现在,我们已经成功地使用Matplotlib设置了坐标轴的纵横比例,使得x和y轴的单位长度相等。我们可以看到图形看起来更加正常,因为没有被拉伸或压缩。
总结起来,我们可以通过设置坐标轴的纵横比例使得x和y轴的单位长度相等。在Matplotlib中,
我们可以使用axis()函数来设置坐标轴的范围和纵横比例,以及使用set_aspect()方法来设置纵横比例。
如果我们将aspect参数设置为"equal",则x和y轴的单位长度将相等。否则,我们可以计算出x和y轴的比例,
并将其作为浮点数提供给aspect参数。